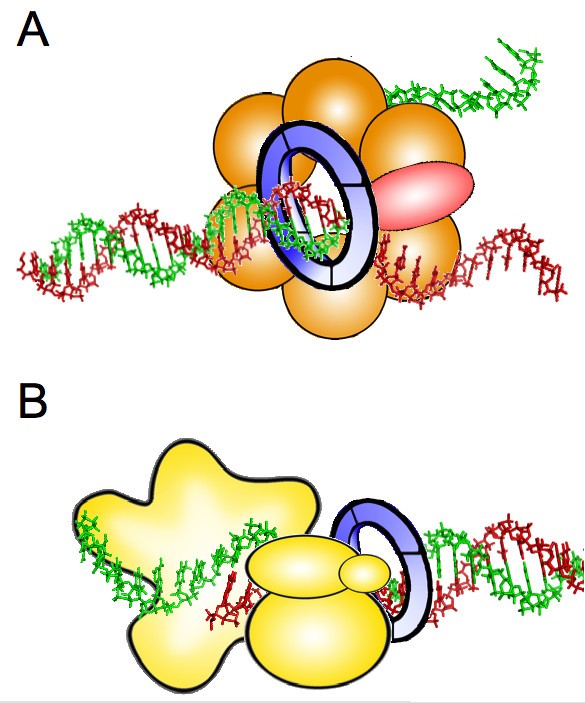
Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation | PNAS
![PDF] The human GINS complex associates with Cdc45 and MCM and is essential for DNA replication | Semantic Scholar PDF] The human GINS complex associates with Cdc45 and MCM and is essential for DNA replication | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2bed3134525627f77020b000c771139c73709545/8-Figure7-1.png)
PDF] The human GINS complex associates with Cdc45 and MCM and is essential for DNA replication | Semantic Scholar
Structural components of the replisome. The CMG replicative helicase... | Download Scientific Diagram

Cdc48 and a ubiquitin ligase drive disassembly of the CMG helicase at the end of DNA replication | Science

CMG helicase and DNA polymerase ε form a functional 15-subunit holoenzyme for eukaryotic leading-strand DNA replication | PNAS

CMG helicase and DNA polymerase ε form a functional 15-subunit holoenzyme for eukaryotic leading-strand DNA replication | PNAS
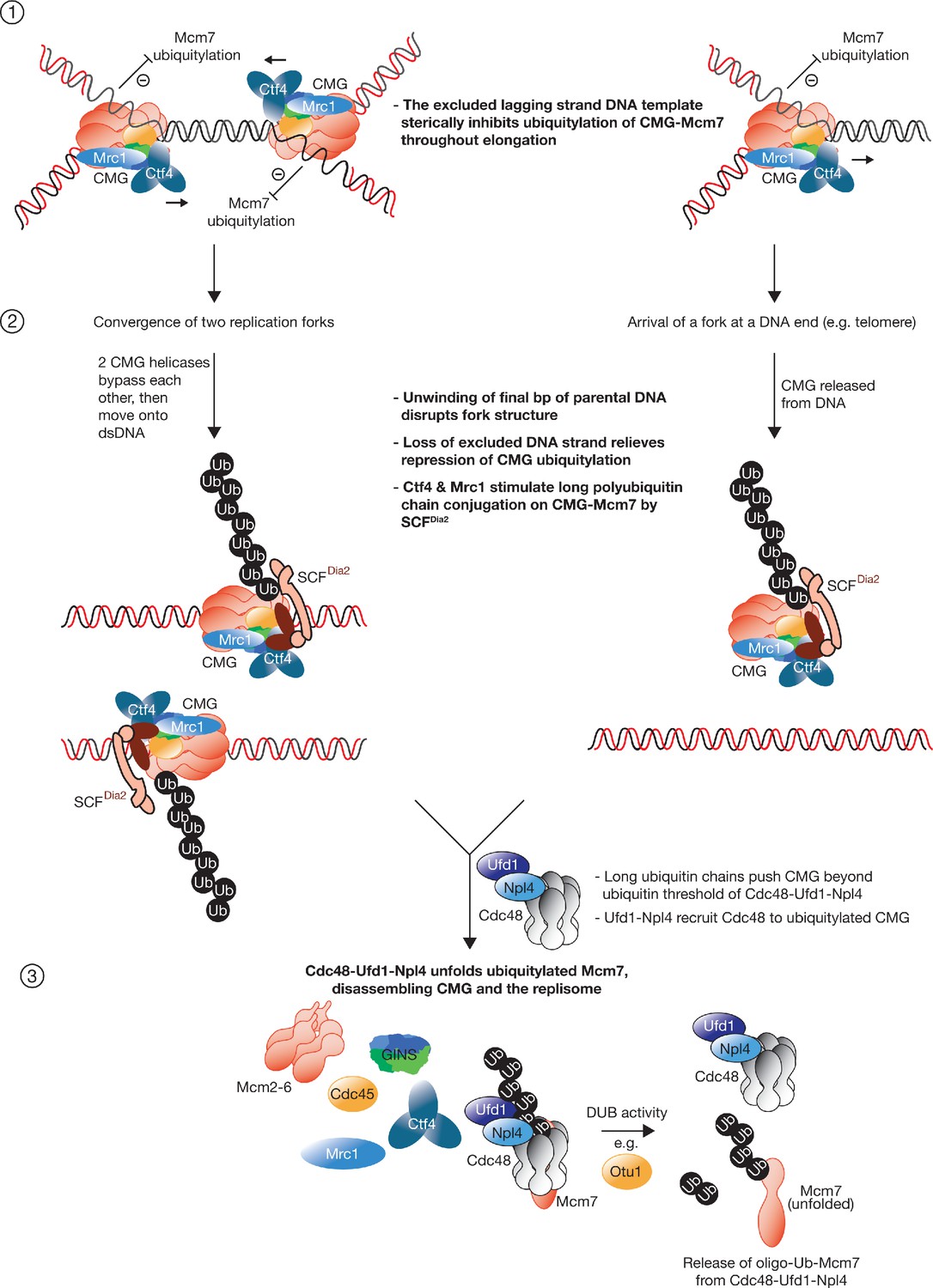
CMG helicase disassembly is controlled by replication fork DNA, replisome components and a ubiquitin threshold | eLife

Mechanisms in helicase activation and analysis of the replication fork activity and architecture – Speck Lab / DNA Replication Group – Christian Speck
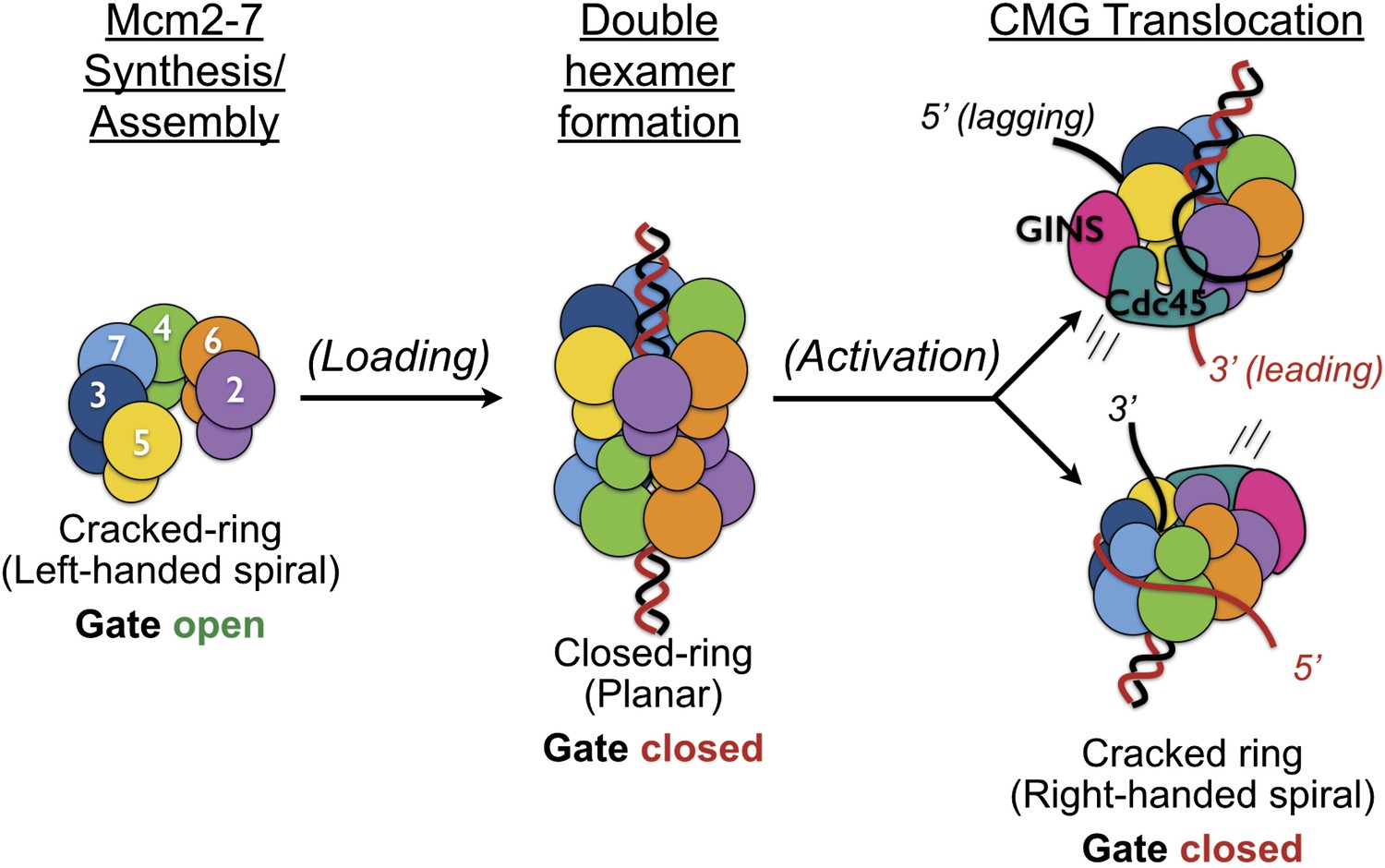
DNA binding polarity, dimerization, and ATPase ring remodeling in the CMG helicase of the eukaryotic replisome | eLife
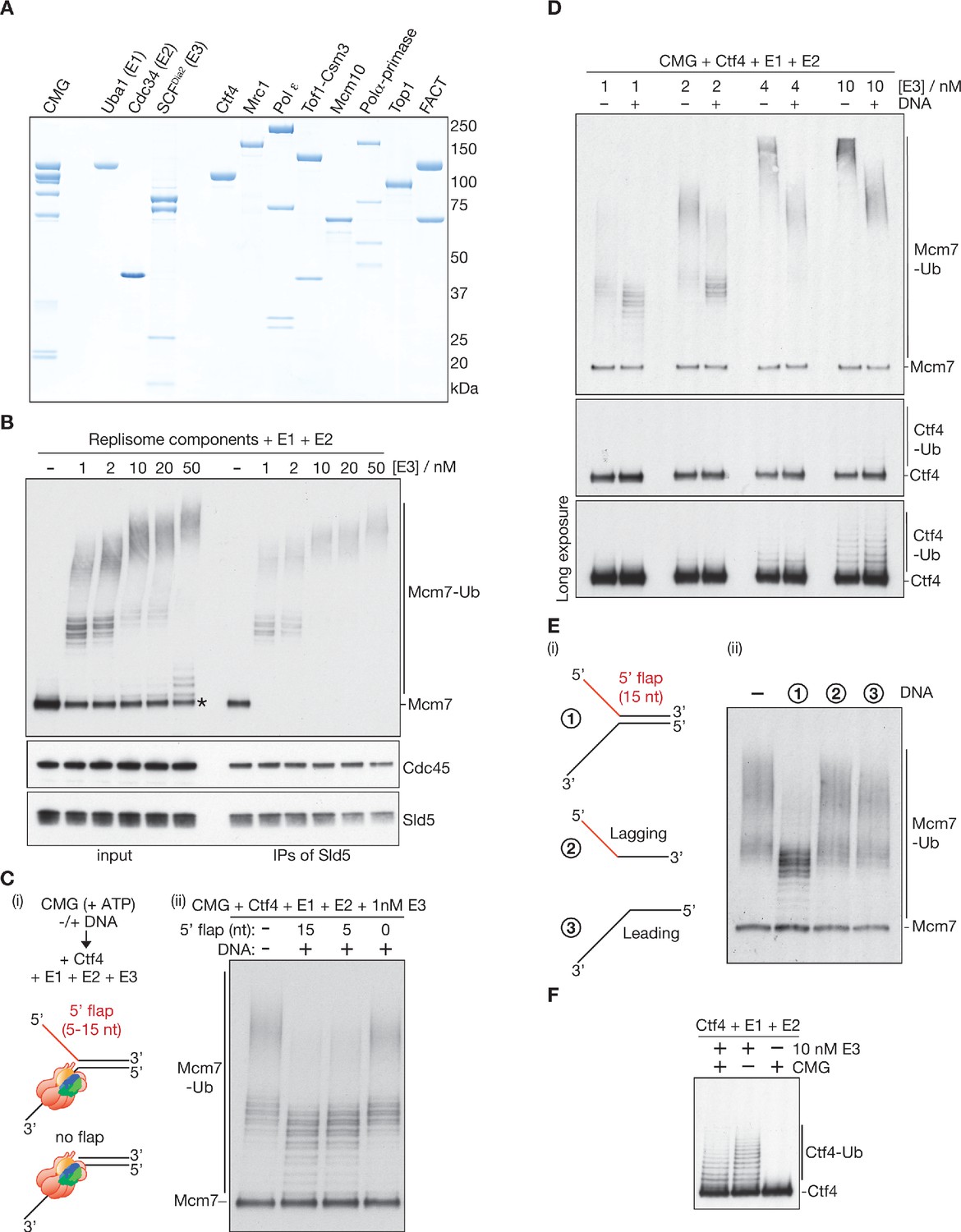
CMG helicase disassembly is controlled by replication fork DNA, replisome components and a ubiquitin threshold | eLife
Isolation of the Cdc45 Mcm2–7 GINS (CMG) complex, a candidate for the eukaryotic DNA replication fork helicase
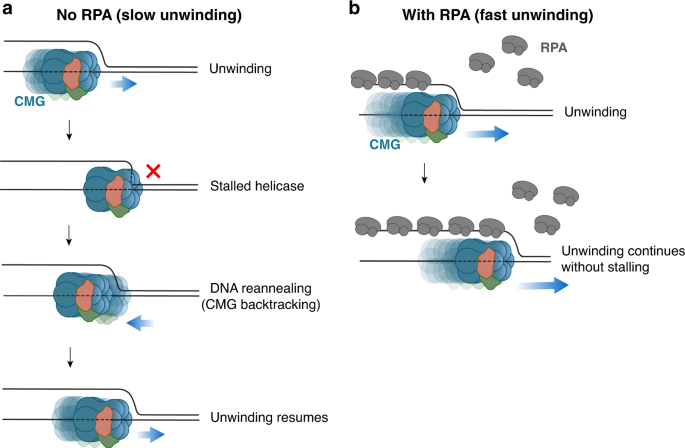
Duplex DNA engagement and RPA oppositely regulate the DNA-unwinding rate of CMG helicase | Nature Communications
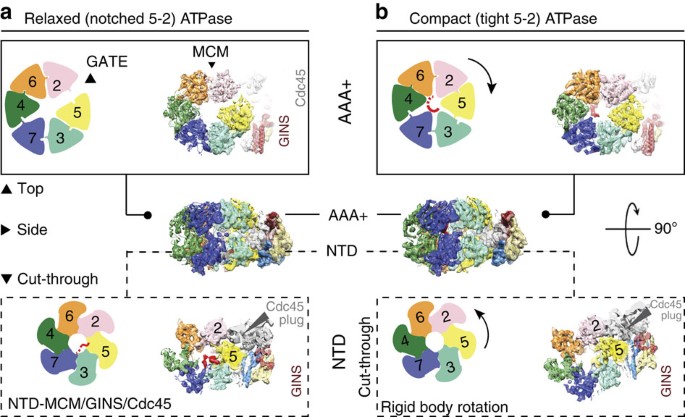
Cryo-EM structures of the eukaryotic replicative helicase bound to a translocation substrate | Nature Communications

3 Summary model of CMG loading and activation during DNA replication.... | Download Scientific Diagram

Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation | PNAS

Mechanistic insights into how CMG helicase facilitates replication past DNA roadblocks - ScienceDirect

DNA densities and interactions with CMG in the CMG-ssDNA structure. CMG... | Download Scientific Diagram

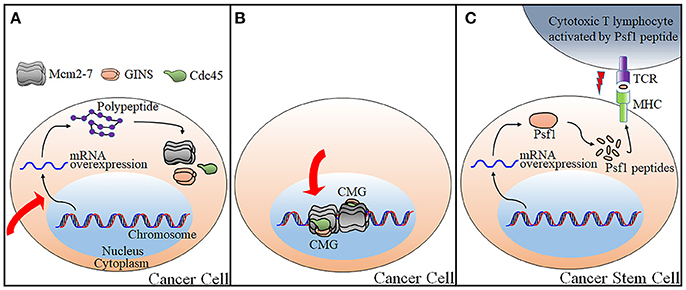
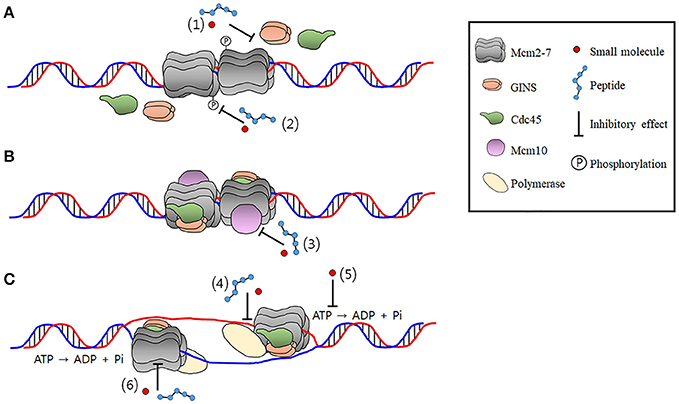
Myc and the Replicative CMG Helicase: The Creation and Destruction of Cancer - Reed - 2020 - BioEssays - Wiley Online Library